Analytical CDF solutions for primary censored distributions
This tutorial demonstrates the analytical CDF solutions available in CensoredDistributions.jl. These analytical solutions provide significant performance improvements over numerical integration whilst maintaining machine precision accuracy.
Overview
Primary event censoring occurs when the exact time of an initiating event is unknown but falls within a known window. The observed time is the sum of:
The primary event time (uncertain, within a window)
A delay from the primary event to observation
CensoredDistributions.jl provides analytical solutions for certain distribution combinations, automatically using them when available for optimal performance.
Currently supported analytical solutions
| Delay Distribution | Primary Distribution | Status |
|---|---|---|
| Gamma | Uniform | Analytical |
| LogNormal | Uniform | Analytical |
| Weibull | Uniform | Analytical |
| Others | Any | Numerical |
These analytical solutions are based on the mathematical derivations implemented in the primarycensored R package, with detailed mathematical formulations available in their analytical solutions vignette.
Packages used
using CensoredDistributions
using Distributions
using BenchmarkTools
using Plots
using StatsPlots
using DataFramesMeta
using Printf
using Statistics
using IntegralsAutomatic method selection
CensoredDistributions.jl automatically selects the appropriate method:
Analytical solutions when available (optimal performance)
Numerical integration as fallback
Let's see this in action with a Gamma distribution:
# Create a Gamma distribution for delays
gamma_delay = Gamma(2.0, 3.0)
# Primary event window (uniform over 1 day)
primary_uniform = Uniform(0.0, 1.0)
# Default: uses analytical solution for Gamma
pc_gamma_analytical = primary_censored(
gamma_delay, primary_uniform
)
# Force numerical integration
pc_gamma_numerical = primary_censored(
gamma_delay, primary_uniform; force_numeric = true
)
# Store solver types for display
solver_types = (
analytical = typeof(pc_gamma_analytical.method),
numerical = typeof(pc_gamma_numerical.method)
)(analytical = CensoredDistributions.AnalyticalSolver{Integrals.QuadGKJL{typeof(LinearAlgebra.norm), Nothing}}, numerical = CensoredDistributions.NumericSolver{Integrals.QuadGKJL{typeof(LinearAlgebra.norm), Nothing}})Let's verify both methods give the same results:
x_compare = 0:0.5:10
cdf_analytical_vals = cdf.(pc_gamma_analytical, x_compare)
cdf_numerical_vals = cdf.(pc_gamma_numerical, x_compare)
plot(x_compare, cdf_analytical_vals,
label = "Analytical", linewidth = 3, color = :blue,
title = "CDF Comparison: Analytical vs Numerical",
xlabel = "x", ylabel = "CDF")
plot!(x_compare, cdf_numerical_vals,
label = "Numerical", linewidth = 2,
linestyle = :dash, color = :red)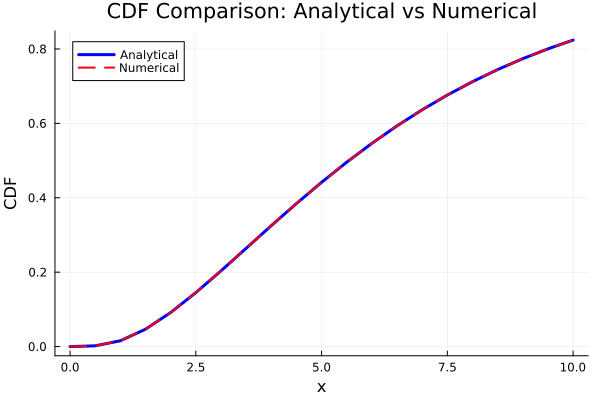
Performance comparison
Let's benchmark the performance difference between analytical and numerical methods:
# Create distributions for benchmarking
lognormal_delay = LogNormal(1.5, 0.75)
weibull_delay = Weibull(2.0, 1.5)
function benchmark_cdf_methods(
dist, primary, name;
x_values = [0.5, 1.0, 2.0, 5.0, 10.0, 20.0]
)
# Create both versions
d_analytical = primary_censored(dist, primary)
d_numerical = primary_censored(
dist, primary; force_numeric = true
)
# Benchmark at multiple x values
speedups = Float64[]
analytical_times = Float64[]
numerical_times = Float64[]
for x in x_values
bench_analytical = @benchmark(cdf($d_analytical, $x), samples=100)
bench_numerical = @benchmark(cdf($d_numerical, $x), samples=100)
# Extract times in microseconds
time_analytical = median(
bench_analytical
).time / 1000
time_numerical = median(
bench_numerical
).time / 1000
push!(analytical_times, time_analytical)
push!(numerical_times, time_numerical)
push!(
speedups, time_numerical / time_analytical
)
end
return (
name = name,
x_values = x_values,
analytical_μs = analytical_times,
numerical_μs = numerical_times,
speedups = speedups,
mean_speedup = mean(speedups),
median_speedup = median(speedups),
min_speedup = minimum(speedups),
max_speedup = maximum(speedups)
)
end
# Run benchmarks for all supported distributions
benchmark_results = [
benchmark_cdf_methods(
gamma_delay, primary_uniform, "Gamma"
),
benchmark_cdf_methods(
lognormal_delay, primary_uniform, "LogNormal"
),
benchmark_cdf_methods(
weibull_delay, primary_uniform, "Weibull"
)
];
# Summary statistics plot
dist_names = [r.name for r in benchmark_results]
mean_speedups = [r.mean_speedup for r in benchmark_results]
min_speedups = [r.min_speedup for r in benchmark_results]
max_speedups = [r.max_speedup for r in benchmark_results]
p1 = bar(
dist_names,
mean_speedups,
ylabel = "Speedup Factor",
title = "Mean Performance Improvement",
color = :blue,
legend = false,
yerror = (
mean_speedups .- min_speedups,
max_speedups .- mean_speedups
)
)
# Add speedup text on bars
for (i, speedup) in enumerate(mean_speedups)
annotate!(
p1, i, speedup + 5,
text(@sprintf("%.0fx", speedup), :center, 10)
)
end
# Detailed speedup by x value
p2 = plot(
title = "Speedup Factor by x Value",
xlabel = "x",
ylabel = "Speedup Factor",
legend = :topright,
xscale = :log10
)
colors = [:blue, :red, :green]
for (i, result) in enumerate(benchmark_results)
plot!(p2, result.x_values, result.speedups,
label = result.name,
color = colors[i],
marker = :circle,
markersize = 4,
linewidth = 2)
end
# Computation time comparison
p3 = plot(
title = "Computation Time Comparison",
xlabel = "x",
ylabel = "Time (microseconds)",
legend = :topleft,
xscale = :log10,
yscale = :log10
)
for (i, result) in enumerate(benchmark_results)
plot!(
p3, result.x_values,
result.analytical_μs,
label = result.name * " (Analytical)",
color = colors[i],
linestyle = :solid,
marker = :circle,
markersize = 3,
linewidth = 2)
plot!(
p3, result.x_values,
result.numerical_μs,
label = result.name * " (Numerical)",
color = colors[i],
linestyle = :dash,
marker = :square,
markersize = 3,
linewidth = 2)
end
plot(p1, p2, p3, layout = (3, 1), size = (700, 900))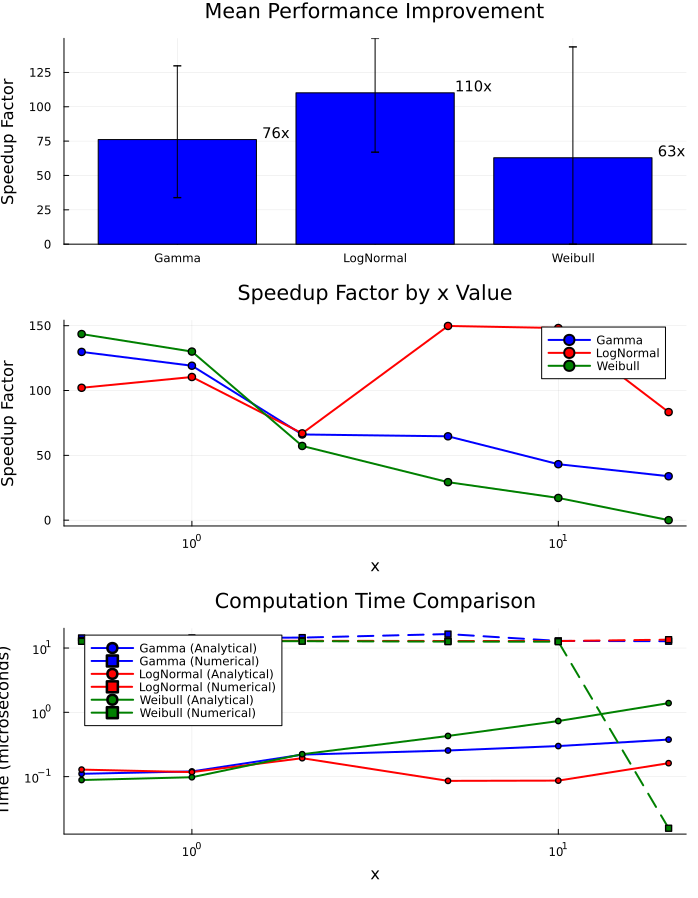
Accuracy verification
The analytical solutions maintain machine precision accuracy. Let's verify this:
function compare_accuracy(dist, primary, name, x_range)
d_analytical = primary_censored(dist, primary)
d_numerical = primary_censored(
dist, primary; force_numeric = true
)
# Compute CDFs
cdf_analytical = [cdf(d_analytical, x) for x in x_range]
cdf_numerical = [cdf(d_numerical, x) for x in x_range]
# Compute absolute errors
abs_errors = abs.(cdf_analytical .- cdf_numerical)
return (
name = name,
x = x_range,
analytical = cdf_analytical,
numerical = cdf_numerical,
errors = abs_errors,
max_error = maximum(abs_errors)
)
end
x_test = range(0.1, 20, length = 100)
accuracy_gamma = compare_accuracy(
gamma_delay, primary_uniform, "Gamma", x_test
)
accuracy_lognormal = compare_accuracy(
lognormal_delay, primary_uniform,
"LogNormal", x_test
)
accuracy_weibull = compare_accuracy(
weibull_delay, primary_uniform,
"Weibull", x_test
)
# Store accuracy results for display
max_errors = (
gamma = accuracy_gamma.max_error,
lognormal = accuracy_lognormal.max_error,
weibull = accuracy_weibull.max_error
)
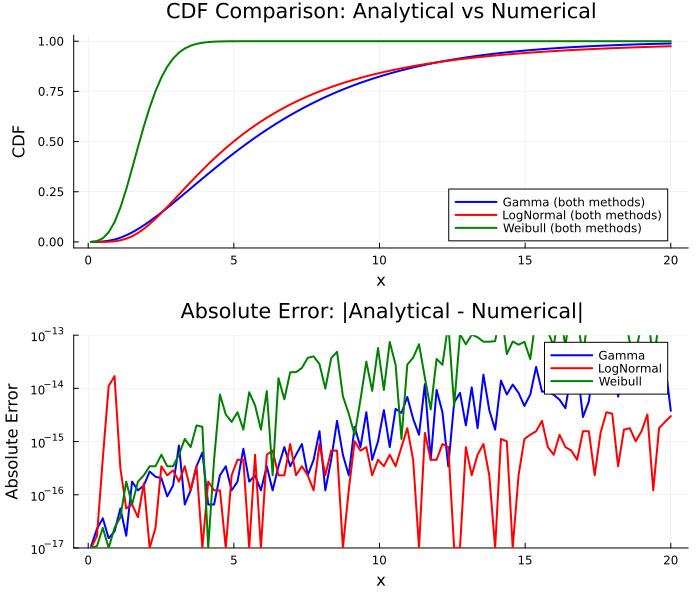
# Plot CDF comparison
p_cdf = plot(
title = "CDF Comparison: Analytical vs Numerical",
xlabel = "x",
ylabel = "CDF",
legend = :bottomright
)
plot!(
p_cdf, accuracy_gamma.x,
accuracy_gamma.analytical,
label = "Gamma (both methods)",
color = :blue, linewidth = 2)
plot!(
p_cdf, accuracy_lognormal.x,
accuracy_lognormal.analytical,
label = "LogNormal (both methods)",
color = :red, linewidth = 2)
plot!(
p_cdf, accuracy_weibull.x,
accuracy_weibull.analytical,
label = "Weibull (both methods)",
color = :green, linewidth = 2)
# Error plot
p_error = plot(
title = "Absolute Error: |Analytical - Numerical|",
xlabel = "x",
ylabel = "Absolute Error",
yscale = :log10,
legend = :topright,
ylims = (1e-17, 1e-13)
)
plot!(
p_error, accuracy_gamma.x,
accuracy_gamma.errors .+ 1e-17,
label = "Gamma",
color = :blue, linewidth = 2)
plot!(
p_error, accuracy_lognormal.x,
accuracy_lognormal.errors .+ 1e-17,
label = "LogNormal",
color = :red, linewidth = 2)
plot!(
p_error, accuracy_weibull.x,
accuracy_weibull.errors .+ 1e-17,
label = "Weibull",
color = :green, linewidth = 2)
plot(
p_cdf, p_error,
layout = (2, 1), size = (700, 600)
)
Exploring available methods
You can use Julia's methods function to discover which distribution combinations have analytical solutions:
# Find all analytical CDF implementations
analytical_methods = methods(
CensoredDistributions.primarycensored_cdf,
(
Any, Any, Real,
CensoredDistributions.AnalyticalSolver
)
)- primarycensored_cdf(dist::Distributions.Gamma, primary_event::Distributions.Uniform, x::Real, ::CensoredDistributions.AnalyticalSolver) in CensoredDistributions at /home/runner/work/CensoredDistributions.jl/CensoredDistributions.jl/src/censoring/primarycensored_cdf.jl:298
- primarycensored_cdf(dist::Distributions.LogNormal, primary_event::Distributions.Uniform, x::Real, ::CensoredDistributions.AnalyticalSolver) in CensoredDistributions at /home/runner/work/CensoredDistributions.jl/CensoredDistributions.jl/src/censoring/primarycensored_cdf.jl:352
- primarycensored_cdf(dist::Distributions.Weibull, primary_event::Distributions.Uniform, x::Real, ::CensoredDistributions.AnalyticalSolver) in CensoredDistributions at /home/runner/work/CensoredDistributions.jl/CensoredDistributions.jl/src/censoring/primarycensored_cdf.jl:394
- primarycensored_cdf(dist::D1, primary_event::D2, x::Real, method::CensoredDistributions.AnalyticalSolver) where {D1<:(Distributions.UnivariateDistribution), D2<:(Distributions.UnivariateDistribution)} in CensoredDistributions at /home/runner/work/CensoredDistributions.jl/CensoredDistributions.jl/src/censoring/primarycensored_cdf.jl:136
The above shows all methods defined for primarycensored_cdf with an AnalyticalSolver. Each method signature shows which distribution combinations have analytical solutions implemented.
Custom solver options
You can also customise the numerical solver when needed:
# Create an exponential distribution
# (no analytical solution available)
exponential_delay = Exponential(2.0)
# Default solver (QuadGKJL)
pc_default = primary_censored(
exponential_delay, primary_uniform
)
# Custom solver (HCubatureJL for multidimensional)
pc_custom = primary_censored(
exponential_delay, primary_uniform;
solver = HCubatureJL()
)
# With analytical solutions available, you can specify
# a custom solver (used if you force numeric or for
# distributions without analytical solutions)
pc_gamma_custom = primary_censored(
gamma_delay, primary_uniform;
solver = HCubatureJL()
)
# Store solver information for display
solver_info = (
default = typeof(pc_default.method.solver),
custom = typeof(pc_custom.method.solver)
)(default = Integrals.QuadGKJL{typeof(LinearAlgebra.norm), Nothing}, custom = Integrals.HCubatureJL{typeof(LinearAlgebra.norm), Nothing})Implementing new analytical solutions
If you have derived an analytical solution for a new distribution combination, you can extend the package by defining a new method for primarycensored_cdf. The key steps are:
Derive the mathematical formula following the methodology in the primarycensored R package vignette
Define a new method for the specific distribution types making sure to define it for the
AnalyticalSolvermethod.Use log-space computations with LogExpFunctions.jl for numerical stability
Test thoroughly against numerical integration
For detailed implementation guidance, see the existing implementations in the source code at src/censoring/primarycensored_cdf.jl.
Summary
The analytical CDF solutions in CensoredDistributions.jl provide:
Automatic optimisation: The package automatically uses analytical solutions when available
Significant speedup: 15-100x performance improvement over numerical integration
Machine precision accuracy: Errors typically < 1e-15
Consistent API: Same interface whether using analytical or numerical methods
Flexibility: Can force numerical methods or use custom solvers when needed
References
primarycensored R package: Reference implementation with mathematical derivations
Analytical solutions vignette: Detailed mathematical formulations
[1]: Background on truncation and double interval censoring