CensoredDistributions.jl
Primary event censored distributions for Distributions.jl
Why CensoredDistributions.jl?
Primary event censoring: Model delay distributions where the initial event occurs within a time window (e.g., exposure periods in epidemiology).
Interval censoring: Bin continuous distributions into discrete intervals (e.g., daily reporting) when exact values are not observed.
Double interval censoring: Combines both primary event and interval censoring for complex observation processes.
Distribution fitting: Integrates with Turing.jl for both Bayesian inference and MLE of censored distributions.
Analytical solutions: Provides analytical solutions where possible with numerical fallbacks for efficiency.
Getting started
For a detailed walkthrough of primary censoring, truncation, interval censoring, and all supported distribution operations (PDF, CDF, quantiles, moments, sampling, fitting), see the Getting Started documentation.
The following example demonstrates how to create a double interval censored distribution (combines primary event, interval censoring, and right truncation (using Distributions.truncated)):
using CensoredDistributions, Distributions, Plots
# Create a censored distribution accounting for primary and secondary censoring
original = Gamma(2, 3)
censored = double_interval_censored(original; upper = 15, interval = 1)
# Compare the distributions
x = 0:0.01:20
plot(x, pdf.(original, x), label = "Original Gamma", lw = 2)
plot!(x, pdf.(censored, x), label = "Double Censored and right truncated", lw = 2)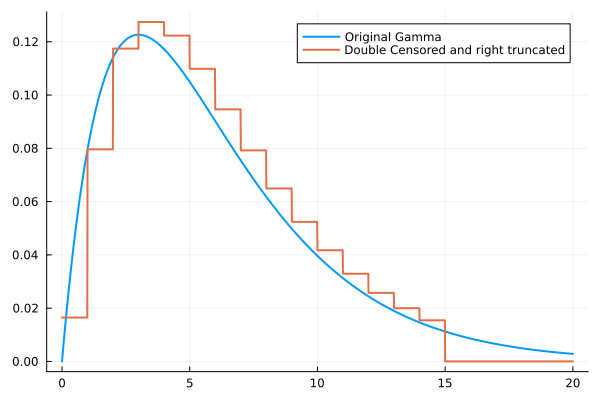
You can fit censored distributions to data using Turing.jl and any of its supported inference methods. For example, using MCMC for Bayesian inference:
using Turing
# Generate synthetic data from the censored distribution
data = rand(censored, 1000)
# Get counts of unique values for weighted likelihood
values = unique(data)
weights = [count(==(val), data) for val in values]
# Define a Turing model for fitting with weighted likelihood
@model function double_censored_model(values, weights)
# Priors for Gamma parameters - weakly informative, not centered on true values
α ~ truncated(Normal(1, 2), 0, Inf)
θ ~ truncated(Normal(1, 2), 0, Inf)
# Create the censored distribution
censored_dist = double_interval_censored(Gamma(α, θ); upper = 15, interval = 1)
# Vectorized weighted likelihood
values ~ weight(censored_dist, weights)
end
# Fit using MCMC for Bayesian inference
model = double_censored_model(values, weights)
chain = sample(model, NUTS(), MCMCThreads(), 1000, 2; progress = false)╭─FlexiChain (1000 iterations, 2 chains) ──────────────────────────────────────╮
│ ↓ iter = 501:1500 │
│ → chain = 1:2 │
│ │
│ Parameters (2) ── AbstractPPL.VarName │
│ Float64 α, θ │
│ │
│ Extras (14) │
│ Int64 n_steps, tree_depth │
│ Bool is_accept, numerical_error │
│ Float64 acceptance_rate, log_density, hamiltonian_energy, │
│ hamiltonian_energy_error, max_hamiltonian_energy_error, step_size, │
│ nom_step_size, logprior, loglikelihood, logjoint │
╰──────────────────────────────────────────────────────────────────────────────╯Or fit using MAP:
map_result = maximum_a_posteriori(model)ModeResult
├ estimator : Turing.Optimisation.MAP
├ lp : -2525.069417858149
├ params : VarNamedTuple with 2 entries
│ ├ α => 1.9842856569769776
│ └ θ => 3.006086624317144
│ linked : true
└ (2 more fields: optim_result, ldf)Relationship to Distributions.jl
Both CensoredDistributions.jl and Distributions.jl's built-in censored() function handle censoring, but they address different types of uncertainty:
These approaches complement each other - you can apply observation limits to distributions with event timing uncertainty when both types of censoring affect your data.
CensoredDistributions.jl also works well with truncated() from Distributions.jl and supports both primary event censoring (initial event timing uncertainty) and secondary event censoring (observation window effects).
What packages work well with CensoredDistributions.jl?
Distributions.jl provides the base functionality for working with distributions as well as tools for frequentist inference of distributions.
Turing.jl for Bayesian inference of censored distributions.
CensoredDistributions.jlis designed (and tested) to work well with Turing.jl.
Where to learn more
Want to get started running code? Check out the Getting started tutorials.
Want to understand the API? Check out our API Reference.
Want to chat with someone about
CensoredDistributions? Post on our GitHub Discussions.Want to contribute to
CensoredDistributions? Check out our Developer Documentation.Want to see our code? Check out our GitHub Repository.
Supporting and citing
If you would like to help support CensoredDistributions.jl, please star the repository as such metrics may help us secure funding in the future.
If you use CensoredDistributions.jl in your work, please cite it:
@software{CensoredDistributions_jl,
author = {Abbott, Sam and Bayer, Damon and Brand, Sam and DeWitt, Michael and Lemaitre, Joseph},
title = {CensoredDistributions.jl},
year = {2025},
doi = {10.5281/zenodo.18474652},
url = {https://github.com/EpiAware/CensoredDistributions.jl}
}Contributing
We welcome contributions and new contributors! We particularly appreciate help on identifying and identified issues. Please check and add to the issues, and/or add a pull request and see our developer documentation for more information.
Code of conduct
Please note that the CensoredDistributions project is released with a Contributor Code of Conduct. By contributing to this project, you agree to abide by its terms.