How exponentially tilted primary events affect observed delay distributions
Introduction
What are we going to do in this exercise
We'll demonstrate how epidemic growth creates additional bias in observed delay distributions through primary event censoring - distinct from truncation bias. We'll cover:
Exponentially tilted vs Uniform primary events
Double interval censoring effects
Impact on delay distributions
What might I need to know before starting
This tutorial builds on Getting Started with CensoredDistributions.jl and focusses on exponentially tilted primary events - when primary events occur non-uniformly within observation windows (due here to exponential dynamics).
During epidemic growth/decline, primary events don't occur uniformly within our observation window:
Growth phase: Recent primary events over-represented leading to shorter observed delays
Decline phase: Older primary events more represented leading to longer observed delays
Steady state: Uniform timing leading to minimal additional bias
Setup
Packages used
using CensoredDistributions
using Distributions
using Plots
using StatsPlots
using DataFramesMeta
using Statistics
using Random
Random.seed!(123)Random.TaskLocalRNG()Simulated scenarios
We'll examine epidemic phase bias using a realistic scenario:
True incubation period: Gamma(4.0, 1.5) with mean 6.0 days
Primary event windows: 7-day observation periods
Growth rate scenarios: r in
true_delay = Gamma(4.0, 1.5);Part 1: Exponentially tilted vs uniform primary events
First, we compare how exponentially tilted distributions with different growth rates shape primary event timing compared to uniform (steady state) patterns.
window_length = 7
scenarios_df = @chain DataFrame(
name = [
"Decline 10%", "Decline 5%",
"Steady", "Growth 5%", "Growth 10%"
],
r = [-0.10, -0.05, 0.0, 0.05, 0.10],
color = [:darkgreen, :lightgreen, :blue, :orange, :red]
) begin
@transform :window = window_length
end
@chain scenarios_df begin
@transform!(:primary_event=ExponentiallyTilted.(0.0, :window, :r))
@transform!(:uniform_ref=Uniform.(0.0, :window))
end| Row | name | r | color | window | primary_event | uniform_ref |
|---|---|---|---|---|---|---|
| String | Float64 | Symbol | Int64 | Exponent… | Uniform… | |
| 1 | Decline 10% | -0.1 | darkgreen | 7 | ExponentiallyTilted{Float64}(min=0.0, max=7.0, r=-0.1) | Distributions.Uniform{Float64}(a=0.0, b=7.0) |
| 2 | Decline 5% | -0.05 | lightgreen | 7 | ExponentiallyTilted{Float64}(min=0.0, max=7.0, r=-0.05) | Distributions.Uniform{Float64}(a=0.0, b=7.0) |
| 3 | Steady | 0.0 | blue | 7 | ExponentiallyTilted{Float64}(min=0.0, max=7.0, r=0.0) | Distributions.Uniform{Float64}(a=0.0, b=7.0) |
| 4 | Growth 5% | 0.05 | orange | 7 | ExponentiallyTilted{Float64}(min=0.0, max=7.0, r=0.05) | Distributions.Uniform{Float64}(a=0.0, b=7.0) |
| 5 | Growth 10% | 0.1 | red | 7 | ExponentiallyTilted{Float64}(min=0.0, max=7.0, r=0.1) | Distributions.Uniform{Float64}(a=0.0, b=7.0) |
Created 5 epidemic scenarios for comparison. Now lets plot the primary event timing distributions.
x_primary = range(0, window_length, length = 100)
p1 = plot(
title = "Primary Event Timing: " *
"ExponentiallyTilted vs Uniform",
xlabel = "Days before observation end",
ylabel = "Probability density",
size = (700, 400), legend = :topright
)
for row in eachrow(scenarios_df)
y_exponential = pdf.(row.primary_event, x_primary)
plot!(p1, x_primary, y_exponential,
label = "$(row.name) (r=$(row.r))",
color = row.color, linewidth = 3)
end
uniform_ref = Uniform(0.0, window_length)
y_uniform = pdf.(uniform_ref, x_primary)
plot!(p1, x_primary, y_uniform,
label = "Uniform (reference)", color = :black,
linestyle = :dash, linewidth = 2)
p1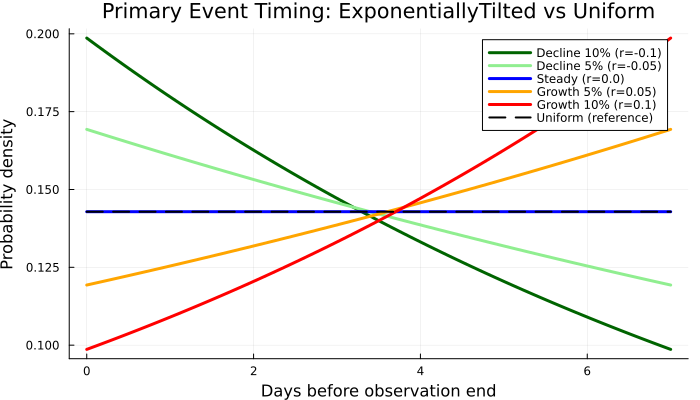
Part 2: Double interval censoring - primary + secondary windows
Now we demonstrate the double interval censoring concept: primary events occur within surveillance-defined primary windows, but during epidemics their timing distribution becomes non-uniform (ExponentiallyTilted), then we observe delays within secondary windows (surveillance window minus primary event time). For the secondary event the distribution doesn't impact what we observe (see references for why).
n_samples = 1000000
secondary_window = 7.0
censoring_windows_df = @chain scenarios_df begin
vcat([DataFrame(
name = fill(row.name, n_samples),
r = fill(row.r, n_samples),
color = fill(row.color, n_samples),
sample_id = 1:n_samples,
primary_time = rand(
row.primary_event, n_samples
)
) for row in eachrow(_)]...)
@transform(:effective_window=secondary_window .- :primary_time)
end;Now we can plot the distribution of effective censoring windows by scenario.
p2 = plot(
title = "Distribution of Effective " *
"Censoring Windows by Epidemic Phase",
xlabel = "Effective censoring window (days)",
ylabel = "Density",
size = (700, 400),
legend = :topright
)
for scenario_name in unique(censoring_windows_df.name)
scenario_data = @subset(censoring_windows_df,
:name .== scenario_name)
scenario_color = scenario_data.color[1]
density!(p2, scenario_data.effective_window,
label = scenario_name,
color = scenario_color,
linewidth = 3,
alpha = 0.7)
end
p2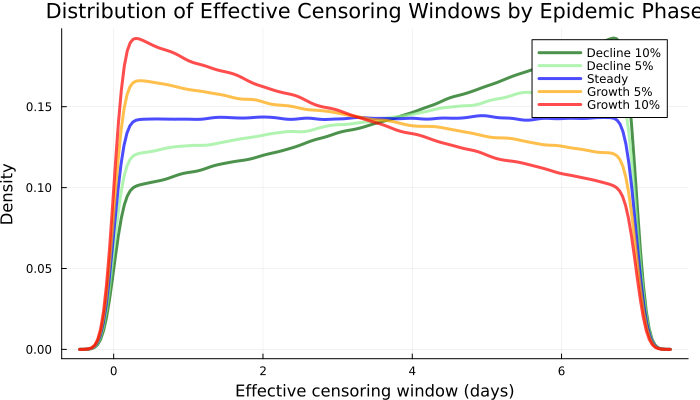
Part 3: Impact on censored delay distributions
Now we examine how epidemic phase bias affects the delays we might observe. We start with the primary censored delay distributions. Note that most of the time, we won't observe these continuous distributions as the secondary event will also be censored.
@chain scenarios_df begin
@transform!(:censored_dist=primary_censored.(
Ref(true_delay), :primary_event
))
end
x_delay = range(0, 15, length = 100)
p3 = plot(
title = "Observed vs True Delay Distributions",
xlabel = "Delay (days)",
ylabel = "Probability density",
size = (700, 400)
)
y_true = pdf.(true_delay, x_delay)
plot!(p3, x_delay, y_true,
label = "True distribution", color = :black,
linestyle = :dash, linewidth = 3)
for row in eachrow(scenarios_df)
y_obs = pdf.(row.censored_dist, x_delay)
plot!(p3, x_delay, y_obs,
label = "$(row.name) (r=$(row.r))",
color = row.color, linewidth = 2)
end
p3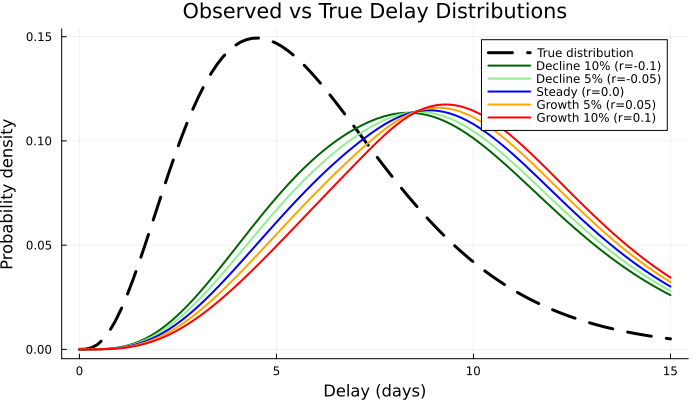
We can also plot the impact on the double censored distribution (which is what we would often observe). Here we show the CDF.
@chain scenarios_df begin
@rtransform!(:double_censored_dist=double_interval_censored(
true_delay;
primary_event = :primary_event,
interval = secondary_window
))
end
x_delay_d = range(0, 40, length = 100)
y_true_d = cdf.(true_delay, x_delay_d)
p4 = plot(
title = "Double vs Single Interval " *
"Censored Distributions",
xlabel = "Delay (days)",
ylabel = "Probability density",
size = (700, 400)
)
plot!(p4, x_delay_d, y_true_d,
label = "True distribution", color = :black,
linestyle = :dash, linewidth = 3)
for row in eachrow(scenarios_df)
y_double = cdf.(
row.double_censored_dist, x_delay_d
)
plot!(p4, x_delay_d, y_double,
label = "$(row.name) Double (r=$(row.r))",
color = row.color, linewidth = 2,
linestyle = :solid)
end
p4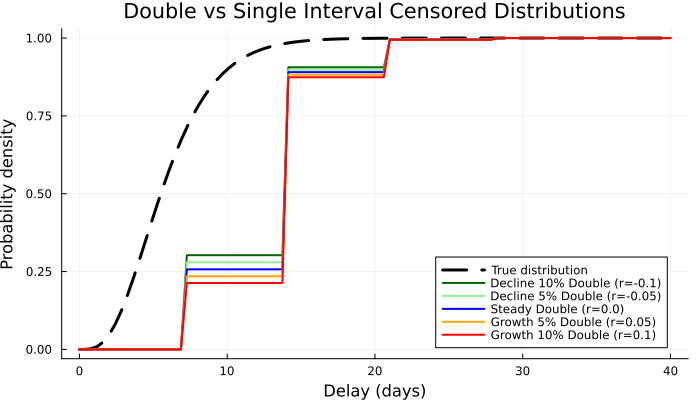
Key insights
- Primary event bias is distinct from truncation bias:
Occurs regardless of the primary event distribution but is more complex when the primary event distribution is non-uniform.
Growth phase: Recent primary events over-represented leading to shorter observed delays
Decline phase: Older primary events more represented leading to longer observed delays
- Two key factors control bias magnitude:
Growth rate magnitude: Stronger growth (larger |r|) creates more divergence from the uniform case
Window length: Longer windows increases divergence for same growth rate
- When to use
ExponentiallyTilted:
Most important when primary censoring intervals are wide (multi-day windows)
For daily primary censoring and moderate epidemic growth, Uniform distributions are often a reasonable approximation
Key benefit of Uniform: Analytical solutions available that are much faster computationally
Use
ExponentiallyTiltedwhen precision is critical, the computational cost is acceptable, and the growth rate is relatively well known.
References
[1]: "Estimating epidemiological delay distributions for infectious diseases"
[2]: "Best practices for estimating and reporting epidemiological delay distributions"
SISMID Tutorial: Interactive bias demonstrations